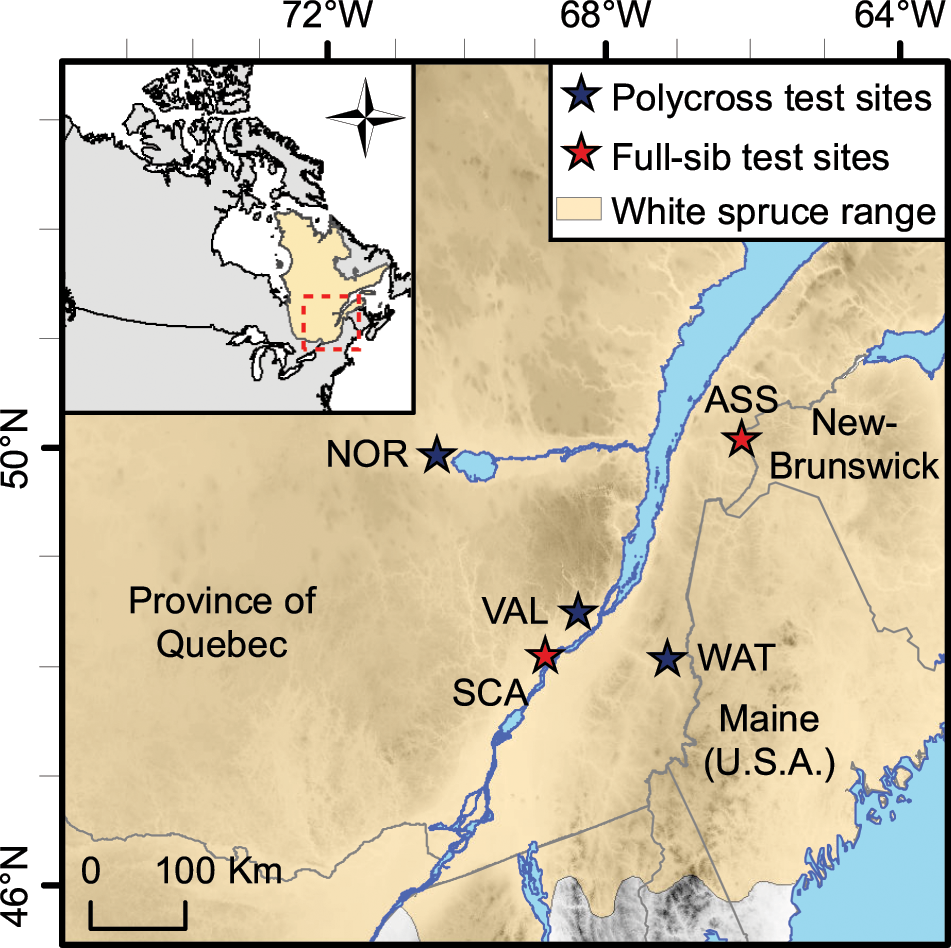
GD between two individuals or populations was defined by Goodman and Lasker (1974) and Nei (1974) as the proportion of non-matching nucleotide bases at homologous nucleotide sites between the genomes of two individuals or two populations. One possible explanation for the limited relationship between pedigree divergence and progeny variance observed in these studies is that coefficients of parentage may be inaccurate because the parents of some specific crosses might differ for many genes affecting a trait.Īn alternative to estimating parental genetic divergence on the basis of pedigree information is the use of molecular marker-based genetic distance (GD). However, other studies in the same species ( Moser and Lee, 1994 Helms et al., 1997 Kisha et al., 1997 Burkhamer et al., 1998 Bohn et al., 1999) indicated that the relationship between parental pedigree distance and progeny genetic variance was neither consistent nor strong enough to permit reliable prediction of genetic variance. Studies in wheat ( Triticum aestivum), oat ( Avena sativa) and soybean ( Glycine max) suggest that pedigree divergence between parents, estimated on the basis of the coefficient of parentage ( Kempthorne, 1969), could be used to predict genetic variance in F 2 or later segregating generations ( Bhatt, 1970, 1973 Cowen and Frey, 1987 Manjarrez-Sandoval et al., 1997).
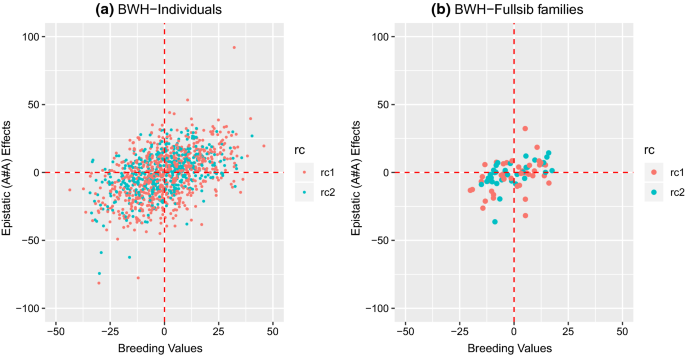
Theoretically, progeny variance is increased in crosses between genetically more distant parents because the number of segregating loci is maximized ( Cox et al., 1984). Maximizing genetic variance results in higher power of QTL detection. Thus, methods that could effectively predict which families are maximally segregating for genetic variation for a trait would aid geneticists by permitting efficient use of resources. Each mapping family is composed of 200 RILs hence, evaluation of the entire population requires testing 5000 inbred lines, beyond the capability of many researchers to assay, particularly for phenotypes that are difficult to measure. For example, the publicly available maize nested association mapping (NAM) population consists of 25 recombinant inbred line (RIL) families derived from crosses between the reference parent B73 and 25 diverse inbred lines ( McMullen et al., 2009). Geneticists have abundant choices of parents to use for mapping population development, and may have numerous extant mapping populations from which to choose for mapping quantitative trait loci (QTL) ( Young, 1996). This architecture, common to many quantitative traits in maize, limits the predictive value of parental genotypic or phenotypic values on progeny variance. These results are congruent with models of genetic architecture that posit numerous genes affecting quantitative traits, each segregating for allelic series, with dispersal of allelic effects across diverse genetic material. Consequently, the choice of promising segregating populations can be based on selecting phenotypically diverse parents. In contrast, we observed for about half of the traits measured a positive correlation between phenotypic parental distances and within-family genetic variance estimates. GDs among parents had no predictive value for progeny variation, which is most likely due to the choice of neutral markers. Genetic distances (GDs) among parents were estimated with 44 simple sequence repeat and 2303 single-nucleotide polymorphism markers. Parents and families were evaluated for 19 quantitative traits across up to 11 environments. We developed 25 families composed of ∼200 random recombinant inbred lines each from crosses between a common reference parent inbred, B73, and 25 diverse maize inbreds. We tested the reliability of genotypic and phenotypic distance estimators between pairs of maize inbred lines to predict genotypic variation for quantitative traits within families derived from biparental crosses. Accurate prediction of the most useful parental combinations within a species would help guide quantitative genetics studies.
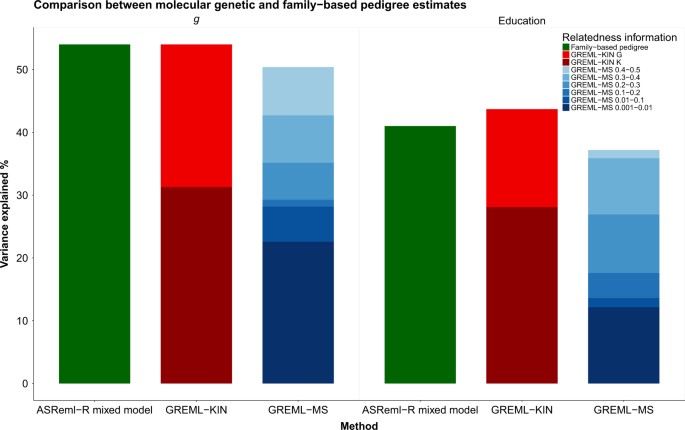
Trait genotypic variation within a family is indicative of the family's informativeness for genetic studies. Appropriate selection of parents for the development of mapping populations is pivotal to maximizing the power of quantitative trait loci detection.
